Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN-132 [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN-132 Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 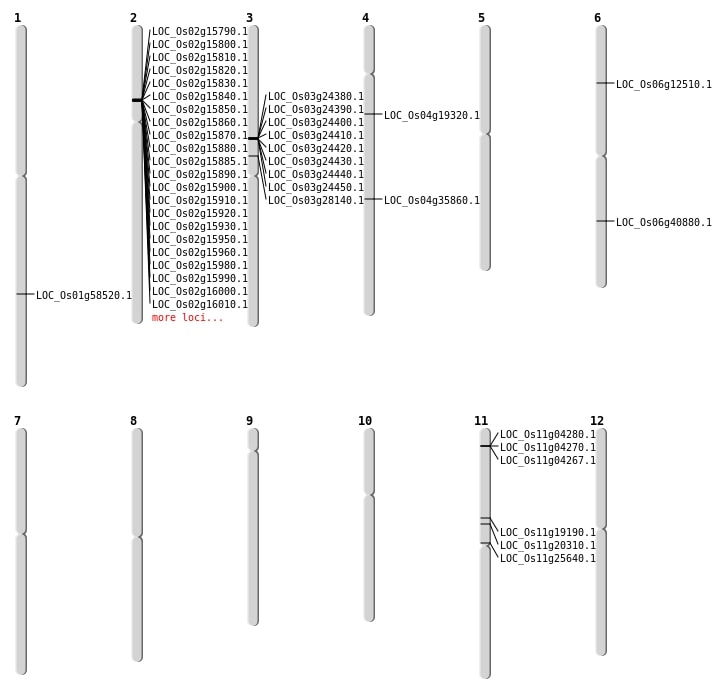
|
| Variant Information |
|---|
Single base substitutions: 34
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 23387307 | A-C | HET | INTERGENIC | |
| SBS | Chr1 | 24209267 | T-A | HET | INTERGENIC | |
| SBS | Chr1 | 24209268 | C-A | HET | INTERGENIC | |
| SBS | Chr10 | 20347630 | G-C | HET | INTERGENIC | |
| SBS | Chr10 | 4022186 | C-T | HET | INTERGENIC | |
| SBS | Chr11 | 14623409 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os11g25640 |
| SBS | Chr11 | 8269516 | G-A | HET | SYNONYMOUS_CODING | |
| SBS | Chr12 | 14304436 | A-T | HOMO | INTERGENIC | |
| SBS | Chr12 | 14672665 | C-A | HOMO | INTERGENIC | |
| SBS | Chr12 | 24604536 | G-A | HET | INTERGENIC | |
| SBS | Chr12 | 3006092 | A-T | HET | INTERGENIC | |
| SBS | Chr2 | 27859241 | C-A | HET | INTERGENIC | |
| SBS | Chr2 | 6068901 | G-A | HOMO | INTERGENIC | |
| SBS | Chr2 | 8186011 | G-T | HET | INTERGENIC | |
| SBS | Chr4 | 10750595 | A-C | HOMO | NON_SYNONYMOUS_CODING | LOC_Os04g19320 |
| SBS | Chr4 | 11046266 | T-C | HOMO | INTERGENIC | |
| SBS | Chr4 | 12319850 | A-G | HOMO | INTERGENIC | |
| SBS | Chr4 | 23800004 | G-A | HET | INTERGENIC | |
| SBS | Chr4 | 25834734 | C-T | HET | UTR_3_PRIME | |
| SBS | Chr4 | 33699207 | A-G | HOMO | INTRON | |
| SBS | Chr4 | 6826323 | G-T | HOMO | INTERGENIC | |
| SBS | Chr5 | 17479434 | G-A | HOMO | INTERGENIC | |
| SBS | Chr6 | 19378138 | T-G | HET | INTERGENIC | |
| SBS | Chr6 | 22498571 | G-A | HOMO | SYNONYMOUS_CODING | |
| SBS | Chr6 | 5434705 | G-T | HOMO | INTERGENIC | |
| SBS | Chr6 | 6791216 | A-G | HOMO | NON_SYNONYMOUS_CODING | LOC_Os06g12510 |
| SBS | Chr7 | 21461588 | T-A | HET | INTERGENIC | |
| SBS | Chr7 | 23402026 | G-A | HET | INTERGENIC | |
| SBS | Chr7 | 23902626 | A-G | HET | INTERGENIC | |
| SBS | Chr7 | 3765836 | C-T | HET | INTRON | |
| SBS | Chr8 | 13740633 | T-G | HET | UTR_3_PRIME | |
| SBS | Chr8 | 5350265 | T-C | HOMO | INTERGENIC | |
| SBS | Chr8 | 6938556 | G-T | HOMO | INTERGENIC | |
| SBS | Chr9 | 10049756 | G-A | HET | INTERGENIC |
Deletions: 23
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr11 | 1750226 | 1762788 | 12562 | 3 |
| Deletion | Chr6 | 3965096 | 3965107 | 11 | |
| Deletion | Chr7 | 4799710 | 4799711 | 1 | |
| Deletion | Chr7 | 7037403 | 7037411 | 8 | |
| Deletion | Chr2 | 8924001 | 9135000 | 211000 | 26 |
| Deletion | Chr10 | 9642696 | 9642703 | 7 | |
| Deletion | Chr9 | 10024230 | 10024236 | 6 | |
| Deletion | Chr9 | 10500328 | 10500330 | 2 | |
| Deletion | Chr9 | 11468564 | 11468571 | 7 | |
| Deletion | Chr9 | 11468568 | 11468575 | 7 | |
| Deletion | Chr6 | 12779532 | 12779534 | 2 | |
| Deletion | Chr3 | 13878001 | 13946000 | 68000 | 9 |
| Deletion | Chr11 | 15500654 | 15500657 | 3 | |
| Deletion | Chr3 | 16186000 | 16186002 | 2 | LOC_Os03g28140 |
| Deletion | Chr2 | 17948443 | 17948445 | 2 | |
| Deletion | Chr9 | 18132477 | 18132482 | 5 | |
| Deletion | Chr9 | 21569188 | 21569201 | 13 | |
| Deletion | Chr4 | 21853896 | 21853899 | 3 | LOC_Os04g35860 |
| Deletion | Chr3 | 23039890 | 23039891 | 1 | |
| Deletion | Chr6 | 24380368 | 24380379 | 11 | LOC_Os06g40880 |
| Deletion | Chr1 | 28465679 | 28465698 | 19 | |
| Deletion | Chr1 | 33817436 | 33817454 | 18 | LOC_Os01g58520 |
| Deletion | Chr1 | 39612180 | 39612185 | 5 |
Insertions: 2
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr1 | 2329880 | 2329881 | 2 | |
| Insertion | Chr8 | 19154939 | 19154939 | 1 |
Inversions: 5
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr2 | 8392965 | 9270152 | LOC_Os02g16290 |
| Inversion | Chr2 | 8907308 | 8922770 | LOC_Os02g15790 |
| Inversion | Chr11 | 10964402 | 11736841 | 2 |
| Inversion | Chr11 | 11563667 | 11736839 | LOC_Os11g20310 |
| Inversion | Chr3 | 13946017 | 14188262 | LOC_Os03g24460 |
No Translocation
