Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN1406-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN1406-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 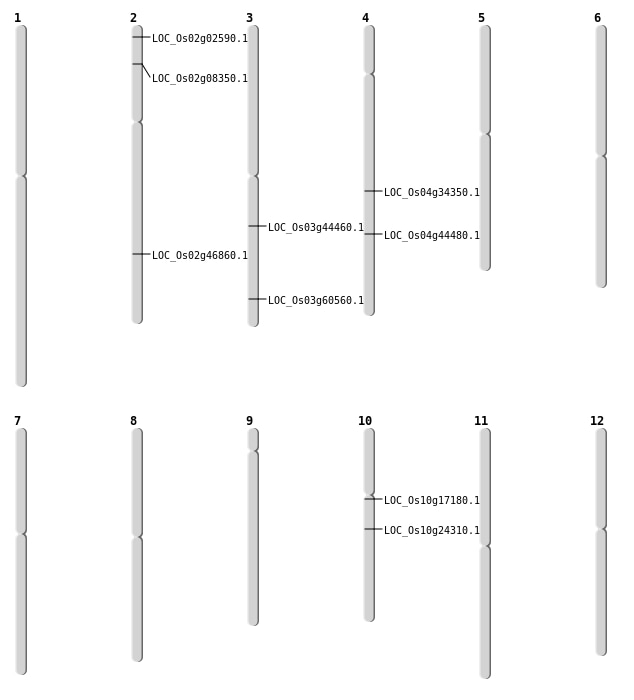
|
| Variant Information |
|---|
Single base substitutions: 27
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 41226515 | A-T | Homo | INTRON | |
| SBS | Chr1 | 42375783 | T-A | Homo | INTERGENIC | |
| SBS | Chr1 | 861459 | C-T | HET | INTERGENIC | |
| SBS | Chr1 | 9487499 | C-T | HET | INTERGENIC | |
| SBS | Chr10 | 8631097 | A-C | HET | NON_SYNONYMOUS_CODING | LOC_Os10g17180 |
| SBS | Chr10 | 9149409 | G-A | HET | INTERGENIC | |
| SBS | Chr2 | 28585489 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os02g46860 |
| SBS | Chr2 | 33777625 | C-T | HET | UTR_5_PRIME | |
| SBS | Chr2 | 7524794 | A-T | HET | INTERGENIC | |
| SBS | Chr2 | 7575693 | T-G | HET | INTERGENIC | |
| SBS | Chr2 | 773648 | C-T | HET | INTERGENIC | |
| SBS | Chr2 | 775365 | T-A | HET | INTERGENIC | |
| SBS | Chr3 | 15497178 | C-T | Homo | INTERGENIC | |
| SBS | Chr3 | 20024076 | C-T | Homo | INTERGENIC | |
| SBS | Chr3 | 25022718 | A-G | HET | NON_SYNONYMOUS_CODING | LOC_Os03g44460 |
| SBS | Chr4 | 18740075 | T-A | HET | INTERGENIC | |
| SBS | Chr4 | 22635577 | G-A | HET | INTERGENIC | |
| SBS | Chr4 | 4809426 | C-T | HET | INTERGENIC | |
| SBS | Chr5 | 17567579 | A-G | Homo | INTRON | |
| SBS | Chr5 | 21470878 | G-A | Homo | INTERGENIC | |
| SBS | Chr6 | 1899399 | G-A | HET | INTERGENIC | |
| SBS | Chr6 | 2607124 | T-A | HET | INTERGENIC | |
| SBS | Chr7 | 21512825 | A-G | HET | INTERGENIC | |
| SBS | Chr7 | 3226839 | A-T | HET | INTRON | |
| SBS | Chr8 | 22811119 | C-T | HET | INTRON | |
| SBS | Chr9 | 11198184 | C-T | HET | SYNONYMOUS_CODING | |
| SBS | Chr9 | 6684193 | C-T | HET | INTERGENIC |
Deletions: 34
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr2 | 938856 | 938857 | 1 | LOC_Os02g02590 |
| Deletion | Chr4 | 1540939 | 1540957 | 18 | |
| Deletion | Chr5 | 2853252 | 2853253 | 1 | |
| Deletion | Chr11 | 4356217 | 4356219 | 2 | |
| Deletion | Chr2 | 4433816 | 4433821 | 5 | LOC_Os02g08350 |
| Deletion | Chr6 | 4608269 | 4608278 | 9 | |
| Deletion | Chr12 | 5011740 | 5011833 | 93 | |
| Deletion | Chr8 | 5093451 | 5093452 | 1 | |
| Deletion | Chr6 | 6919675 | 6919683 | 8 | |
| Deletion | Chr11 | 8610957 | 8610958 | 1 | |
| Deletion | Chr6 | 9658053 | 9658054 | 1 | |
| Deletion | Chr10 | 12453356 | 12453357 | 1 | LOC_Os10g24310 |
| Deletion | Chr1 | 14087096 | 14087097 | 1 | |
| Deletion | Chr9 | 15242109 | 15242112 | 3 | |
| Deletion | Chr8 | 15301500 | 15301501 | 1 | |
| Deletion | Chr10 | 15645626 | 15645631 | 5 | |
| Deletion | Chr10 | 15649225 | 15649229 | 4 | |
| Deletion | Chr5 | 15721767 | 15721769 | 2 | |
| Deletion | Chr2 | 16925612 | 16925613 | 1 | |
| Deletion | Chr9 | 17209479 | 17209481 | 2 | |
| Deletion | Chr4 | 18312356 | 18312357 | 1 | |
| Deletion | Chr1 | 19293225 | 19293234 | 9 | |
| Deletion | Chr4 | 20803850 | 20803857 | 7 | LOC_Os04g34350 |
| Deletion | Chr12 | 22058178 | 22058215 | 37 | |
| Deletion | Chr2 | 22309658 | 22309659 | 1 | |
| Deletion | Chr12 | 22345969 | 22345994 | 25 | |
| Deletion | Chr3 | 22715134 | 22715141 | 7 | |
| Deletion | Chr6 | 22814482 | 22814483 | 1 | |
| Deletion | Chr2 | 23828586 | 23828591 | 5 | |
| Deletion | Chr12 | 25869288 | 25869289 | 1 | |
| Deletion | Chr3 | 26077618 | 26077625 | 7 | |
| Deletion | Chr4 | 26327083 | 26327084 | 1 | LOC_Os04g44480 |
| Deletion | Chr5 | 29396820 | 29396827 | 7 | |
| Deletion | Chr3 | 34424963 | 34424964 | 1 | LOC_Os03g60560 |
Insertions: 18
Inversions: 2
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr7 | 1945295 | 3532793 | |
| Inversion | Chr7 | 3270038 | 3532778 |
No Translocation
