Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN2231-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN2231-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 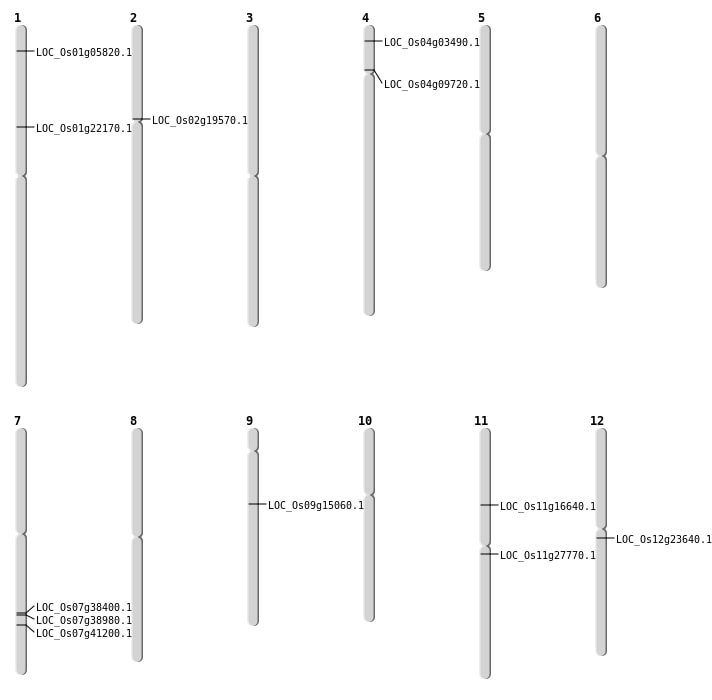
|
| Variant Information |
|---|
Single base substitutions: 48
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 12461868 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os01g22170 |
| SBS | Chr1 | 17116817 | G-A | HET | ||
| SBS | Chr1 | 21169430 | T-G | HET | ||
| SBS | Chr1 | 21169431 | A-G | HET | ||
| SBS | Chr1 | 25115789 | G-A | HET | ||
| SBS | Chr1 | 29240426 | A-T | HET | ||
| SBS | Chr1 | 35328389 | C-T | HET | ||
| SBS | Chr1 | 37656377 | C-G | HET | ||
| SBS | Chr1 | 8180671 | A-G | HET | ||
| SBS | Chr10 | 10419920 | G-A | HET | ||
| SBS | Chr10 | 11256977 | A-G | HET | ||
| SBS | Chr10 | 12427593 | A-G | HET | ||
| SBS | Chr10 | 2985588 | C-T | HET | ||
| SBS | Chr10 | 8818738 | C-T | HET | ||
| SBS | Chr11 | 13746950 | T-C | HET | ||
| SBS | Chr11 | 15988204 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os11g27770 |
| SBS | Chr11 | 24179421 | C-T | HET | ||
| SBS | Chr11 | 4879410 | T-C | HET | ||
| SBS | Chr11 | 9230302 | C-G | HET | NON_SYNONYMOUS_CODING | LOC_Os11g16640 |
| SBS | Chr12 | 13420569 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os12g23640 |
| SBS | Chr12 | 3283881 | A-G | HET | ||
| SBS | Chr2 | 11435831 | C-A | Homo | NON_SYNONYMOUS_CODING | LOC_Os02g19570 |
| SBS | Chr2 | 1198913 | C-A | HET | ||
| SBS | Chr2 | 23152631 | A-G | HET | ||
| SBS | Chr2 | 29096378 | C-T | Homo | ||
| SBS | Chr3 | 12832018 | A-G | HET | ||
| SBS | Chr3 | 21271953 | T-A | HET | ||
| SBS | Chr3 | 25563639 | A-G | HET | ||
| SBS | Chr3 | 33981225 | T-A | HET | ||
| SBS | Chr3 | 35662989 | G-A | HET | ||
| SBS | Chr3 | 35662990 | G-A | HET | ||
| SBS | Chr4 | 12316754 | C-A | Homo | ||
| SBS | Chr4 | 18395644 | C-T | Homo | ||
| SBS | Chr4 | 3145368 | G-A | HET | ||
| SBS | Chr4 | 9810818 | C-T | Homo | ||
| SBS | Chr5 | 12922910 | C-T | Homo | ||
| SBS | Chr5 | 14986169 | G-A | Homo | ||
| SBS | Chr5 | 15506027 | T-A | Homo | ||
| SBS | Chr5 | 25876862 | T-C | Homo | ||
| SBS | Chr6 | 3008456 | T-A | HET | ||
| SBS | Chr7 | 23844233 | A-G | HET | ||
| SBS | Chr7 | 24666409 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os07g41200 |
| SBS | Chr7 | 25219104 | T-C | HET | ||
| SBS | Chr7 | 25632477 | C-T | HET | ||
| SBS | Chr8 | 15153338 | C-T | HET | ||
| SBS | Chr8 | 23084427 | C-T | Homo | ||
| SBS | Chr8 | 5227375 | A-G | HET | ||
| SBS | Chr8 | 8236767 | C-T | HET |
Deletions: 12
Insertions: 8
Inversions: 4
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr9 | 9107148 | 9541174 | LOC_Os09g15060 |
| Inversion | Chr10 | 15585720 | 15657462 | |
| Inversion | Chr10 | 16532244 | 16682271 | |
| Inversion | Chr8 | 16999242 | 17095841 |
Translocations: 14
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr9 | 1100908 | Chr2 | 2674024 | |
| Translocation | Chr4 | 5215507 | Chr1 | 1044499 | LOC_Os04g09720 |
| Translocation | Chr5 | 6371330 | Chr4 | 16078918 | |
| Translocation | Chr10 | 7621619 | Chr5 | 17252612 | |
| Translocation | Chr7 | 8261476 | Chr3 | 18185027 | |
| Translocation | Chr9 | 9107166 | Chr7 | 26204442 | LOC_Os09g15060 |
| Translocation | Chr9 | 9541194 | Chr7 | 26196650 | |
| Translocation | Chr8 | 10828772 | Chr4 | 21079446 | |
| Translocation | Chr11 | 13416165 | Chr4 | 1508364 | 2 |
| Translocation | Chr11 | 15746022 | Chr1 | 2790029 | 2 |
| Translocation | Chr7 | 23081188 | Chr2 | 1820235 | LOC_Os07g38400 |
| Translocation | Chr7 | 23370711 | Chr2 | 19766043 | LOC_Os07g38980 |
| Translocation | Chr11 | 25314483 | Chr8 | 8662999 | |
| Translocation | Chr2 | 27133113 | Chr1 | 25821321 |
