Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN2358 [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN2358 Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 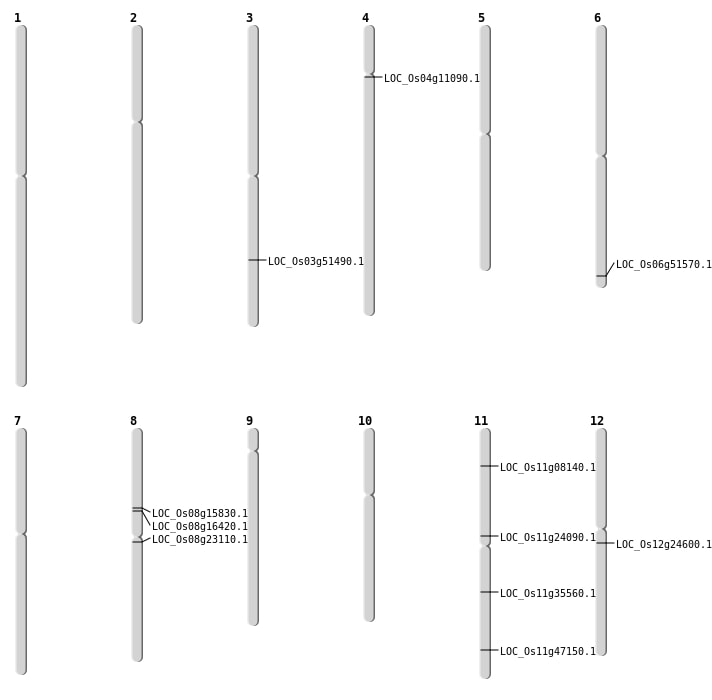
|
| Variant Information |
|---|
Single base substitutions: 45
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 23221842 | T-C | Homo | ||
| SBS | Chr1 | 29959208 | C-G | Homo | ||
| SBS | Chr1 | 3569001 | T-C | HET | ||
| SBS | Chr10 | 14830564 | T-C | HET | ||
| SBS | Chr10 | 19722888 | A-G | HET | ||
| SBS | Chr10 | 9673967 | G-C | Homo | ||
| SBS | Chr11 | 10617712 | A-G | HET | ||
| SBS | Chr11 | 20852412 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os11g35560 |
| SBS | Chr12 | 14076460 | C-G | Homo | NON_SYNONYMOUS_CODING | LOC_Os12g24600 |
| SBS | Chr12 | 19625293 | G-T | Homo | ||
| SBS | Chr2 | 18817884 | G-A | HET | ||
| SBS | Chr2 | 22513503 | T-A | HET | ||
| SBS | Chr2 | 24487923 | C-A | HET | ||
| SBS | Chr2 | 2701643 | A-G | Homo | ||
| SBS | Chr2 | 27935863 | G-A | HET | ||
| SBS | Chr2 | 29670852 | C-A | HET | ||
| SBS | Chr2 | 34060845 | G-A | HET | ||
| SBS | Chr3 | 18108554 | G-A | Homo | ||
| SBS | Chr3 | 23879800 | C-T | Homo | ||
| SBS | Chr3 | 24846797 | T-A | HET | ||
| SBS | Chr3 | 29457365 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os03g51490 |
| SBS | Chr3 | 7038600 | G-A | Homo | ||
| SBS | Chr4 | 11174674 | C-G | HET | ||
| SBS | Chr4 | 27563010 | C-T | HET | ||
| SBS | Chr4 | 34319315 | A-T | HET | ||
| SBS | Chr4 | 34571123 | A-G | HET | ||
| SBS | Chr4 | 6036263 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os04g11090 |
| SBS | Chr5 | 16934224 | G-T | HET | ||
| SBS | Chr5 | 6414104 | G-A | HET | ||
| SBS | Chr6 | 23317818 | T-G | Homo | ||
| SBS | Chr6 | 31243482 | C-T | Homo | NON_SYNONYMOUS_CODING | LOC_Os06g51570 |
| SBS | Chr6 | 7888926 | C-T | HET | ||
| SBS | Chr6 | 8534203 | C-T | HET | ||
| SBS | Chr7 | 17996175 | T-C | HET | ||
| SBS | Chr7 | 23902626 | A-G | HET | ||
| SBS | Chr7 | 24236344 | T-C | HET | ||
| SBS | Chr7 | 5798469 | A-G | HET | ||
| SBS | Chr8 | 13291019 | G-A | HET | ||
| SBS | Chr8 | 13921997 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os08g23110 |
| SBS | Chr8 | 2361810 | G-A | Homo | ||
| SBS | Chr8 | 5912682 | A-T | HET | ||
| SBS | Chr8 | 8420031 | T-G | Homo | ||
| SBS | Chr9 | 11065280 | G-A | HET | ||
| SBS | Chr9 | 2707710 | G-T | Homo | ||
| SBS | Chr9 | 3096355 | C-T | Homo |
Deletions: 1
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr10 | 11763012 | 11763014 | 2 |
Insertions: 1
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr11 | 16685571 | 16685640 | 70 |
No Inversion
Translocations: 7
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr11 | 4272680 | Chr4 | 4022174 | LOC_Os11g08140 |
| Translocation | Chr10 | 5265517 | Chr3 | 13387230 | |
| Translocation | Chr8 | 9624427 | Chr7 | 8806239 | LOC_Os08g15830 |
| Translocation | Chr8 | 9910724 | Chr6 | 11334675 | |
| Translocation | Chr11 | 13696374 | Chr8 | 10037269 | 2 |
| Translocation | Chr3 | 14810910 | Chr2 | 4884867 | |
| Translocation | Chr11 | 28369452 | Chr8 | 2141307 | LOC_Os11g47150 |
