Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN2660-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN2660-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 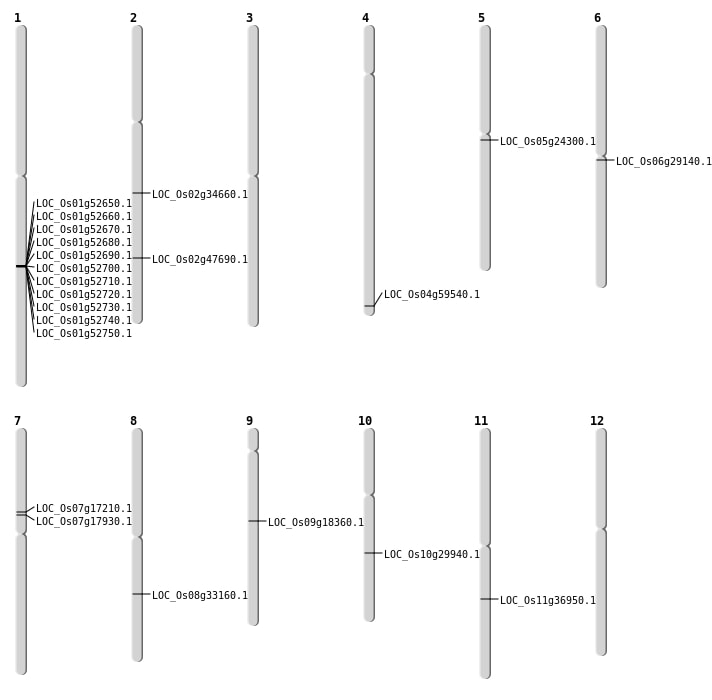
|
| Variant Information |
|---|
Single base substitutions: 51
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 10545776 | C-T | HET | ||
| SBS | Chr1 | 19027873 | G-A | HET | ||
| SBS | Chr1 | 19737218 | T-C | HET | ||
| SBS | Chr1 | 29329544 | T-C | HET | ||
| SBS | Chr1 | 32063713 | T-A | Homo | ||
| SBS | Chr10 | 15545024 | A-G | HET | NON_SYNONYMOUS_CODING | LOC_Os10g29940 |
| SBS | Chr10 | 8726525 | C-T | HET | ||
| SBS | Chr11 | 21798356 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os11g36950 |
| SBS | Chr11 | 24774111 | C-T | HET | ||
| SBS | Chr11 | 25028374 | G-T | HET | ||
| SBS | Chr12 | 10915341 | G-A | HET | ||
| SBS | Chr12 | 14430760 | G-A | HET | ||
| SBS | Chr12 | 14948404 | G-A | HET | ||
| SBS | Chr12 | 15043437 | C-G | HET | ||
| SBS | Chr12 | 248452 | G-A | HET | ||
| SBS | Chr2 | 18885656 | G-A | Homo | ||
| SBS | Chr2 | 20793274 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os02g34660 |
| SBS | Chr2 | 20793276 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os02g34660 |
| SBS | Chr2 | 8355350 | G-A | HET | ||
| SBS | Chr3 | 13326217 | C-A | Homo | ||
| SBS | Chr3 | 16034727 | T-C | HET | ||
| SBS | Chr3 | 16849124 | G-T | HET | ||
| SBS | Chr3 | 18854188 | G-A | HET | ||
| SBS | Chr3 | 23835002 | A-G | HET | ||
| SBS | Chr3 | 28369561 | C-T | HET | ||
| SBS | Chr3 | 4186401 | G-A | HET | ||
| SBS | Chr4 | 12638042 | T-A | Homo | ||
| SBS | Chr4 | 19428291 | C-G | Homo | ||
| SBS | Chr4 | 7420905 | C-T | Homo | ||
| SBS | Chr5 | 14062997 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os05g24300 |
| SBS | Chr5 | 16823315 | C-T | HET | ||
| SBS | Chr5 | 24963282 | T-C | HET | ||
| SBS | Chr5 | 24963283 | A-T | HET | ||
| SBS | Chr5 | 26074346 | G-A | HET | ||
| SBS | Chr7 | 10608441 | G-C | HET | NON_SYNONYMOUS_CODING | LOC_Os07g17930 |
| SBS | Chr7 | 18909743 | C-A | HET | ||
| SBS | Chr7 | 4210481 | C-T | HET | ||
| SBS | Chr7 | 8139444 | T-C | HET | ||
| SBS | Chr8 | 12696462 | T-C | Homo | ||
| SBS | Chr8 | 14518896 | G-T | HET | ||
| SBS | Chr8 | 19492637 | C-T | HET | ||
| SBS | Chr8 | 20626123 | T-G | HET | NON_SYNONYMOUS_CODING | LOC_Os08g33160 |
| SBS | Chr8 | 22564733 | G-T | HET | ||
| SBS | Chr8 | 25750222 | T-C | HET | ||
| SBS | Chr8 | 26581080 | G-A | HET | ||
| SBS | Chr9 | 11260000 | T-C | Homo | NON_SYNONYMOUS_CODING | LOC_Os09g18360 |
| SBS | Chr9 | 11260005 | C-A | Homo | NON_SYNONYMOUS_CODING | LOC_Os09g18360 |
| SBS | Chr9 | 14621063 | G-A | Homo | ||
| SBS | Chr9 | 6591940 | G-A | Homo | ||
| SBS | Chr9 | 6591942 | A-T | Homo | ||
| SBS | Chr9 | 8767262 | C-A | HET |
Deletions: 12
Insertions: 5
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr4 | 14979680 | 14979681 | 2 | |
| Insertion | Chr4 | 2102917 | 2102918 | 2 | |
| Insertion | Chr6 | 16630623 | 16630623 | 1 | LOC_Os06g29140 |
| Insertion | Chr6 | 3255931 | 3255932 | 2 | |
| Insertion | Chr8 | 28224105 | 28224106 | 2 |
No Inversion
Translocations: 13
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr10 | 2491072 | Chr5 | 25142917 | |
| Translocation | Chr10 | 3669339 | Chr4 | 33407149 | |
| Translocation | Chr8 | 4306960 | Chr6 | 25086676 | |
| Translocation | Chr7 | 10136799 | Chr2 | 30553950 | LOC_Os07g17210 |
| Translocation | Chr7 | 10140869 | Chr2 | 30553874 | |
| Translocation | Chr11 | 12007501 | Chr4 | 6643298 | |
| Translocation | Chr7 | 12886311 | Chr5 | 2320280 | |
| Translocation | Chr8 | 13069649 | Chr2 | 9805464 | |
| Translocation | Chr10 | 16821028 | Chr1 | 41587176 | |
| Translocation | Chr12 | 18408964 | Chr4 | 35415968 | 2 |
| Translocation | Chr5 | 18822687 | Chr2 | 29161329 | 2 |
| Translocation | Chr8 | 20440108 | Chr5 | 6354833 | |
| Translocation | Chr7 | 23842203 | Chr1 | 39419953 |
