Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN2685-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN2685-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 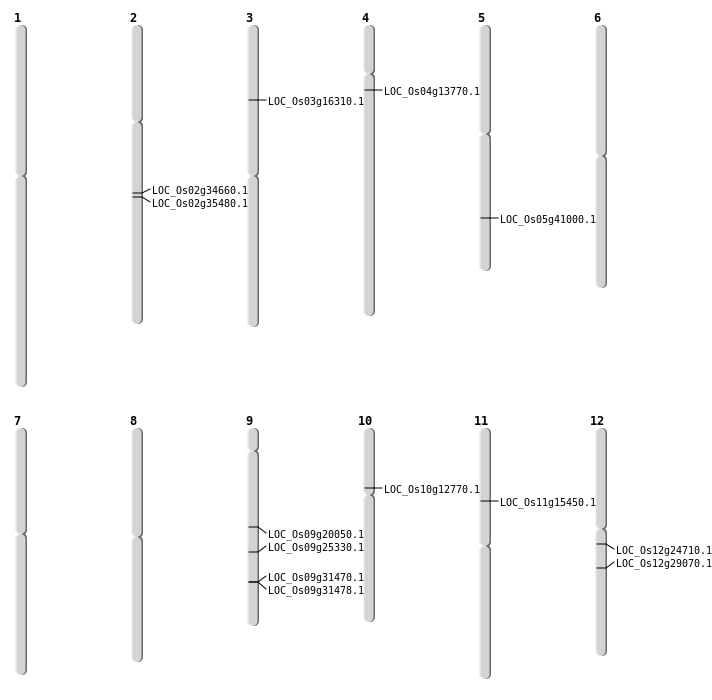
|
| Variant Information |
|---|
Single base substitutions: 37
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 11496653 | C-A | HET | ||
| SBS | Chr1 | 11496655 | G-A | HET | ||
| SBS | Chr1 | 18058726 | A-G | HET | ||
| SBS | Chr1 | 23767168 | T-C | HET | ||
| SBS | Chr1 | 27492239 | C-T | Homo | ||
| SBS | Chr10 | 18930815 | G-A | HET | ||
| SBS | Chr10 | 7133148 | G-A | HET | STOP_GAINED | LOC_Os10g12770 |
| SBS | Chr11 | 10600679 | T-C | Homo | ||
| SBS | Chr11 | 22263715 | C-T | HET | ||
| SBS | Chr12 | 14168660 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os12g24710 |
| SBS | Chr12 | 17220550 | A-G | HET | NON_SYNONYMOUS_CODING | LOC_Os12g29070 |
| SBS | Chr12 | 4397715 | C-T | HET | ||
| SBS | Chr12 | 9371260 | G-C | HET | ||
| SBS | Chr2 | 20214321 | C-T | HET | ||
| SBS | Chr2 | 21326421 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os02g35480 |
| SBS | Chr2 | 24077362 | C-T | HET | ||
| SBS | Chr2 | 24077363 | G-T | HET | ||
| SBS | Chr2 | 32696296 | T-C | HET | ||
| SBS | Chr2 | 4980164 | C-T | HET | ||
| SBS | Chr3 | 12000236 | C-A | HET | ||
| SBS | Chr3 | 33127137 | C-A | Homo | ||
| SBS | Chr3 | 34156870 | G-A | HET | ||
| SBS | Chr3 | 6629974 | G-A | Homo | ||
| SBS | Chr4 | 20778419 | C-T | HET | ||
| SBS | Chr4 | 3074304 | A-T | Homo | ||
| SBS | Chr5 | 24038382 | C-T | Homo | NON_SYNONYMOUS_CODING | LOC_Os05g41000 |
| SBS | Chr5 | 27868291 | A-G | HET | ||
| SBS | Chr5 | 8921660 | C-T | HET | ||
| SBS | Chr6 | 11600107 | A-T | Homo | ||
| SBS | Chr7 | 14982401 | C-T | HET | ||
| SBS | Chr7 | 8021846 | C-G | HET | ||
| SBS | Chr8 | 22233290 | A-T | Homo | ||
| SBS | Chr8 | 3371451 | C-A | HET | ||
| SBS | Chr8 | 9851255 | A-T | Homo | ||
| SBS | Chr9 | 12017827 | A-T | Homo | NON_SYNONYMOUS_CODING | LOC_Os09g20050 |
| SBS | Chr9 | 15163890 | G-A | Homo | NON_SYNONYMOUS_CODING | LOC_Os09g25330 |
| SBS | Chr9 | 15920521 | A-G | HET |
Deletions: 10
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr12 | 5306851 | 5306870 | 19 | |
| Deletion | Chr11 | 8758044 | 8758049 | 5 | LOC_Os11g15450 |
| Deletion | Chr3 | 9006162 | 9006167 | 5 | LOC_Os03g16310 |
| Deletion | Chr11 | 11799161 | 11799163 | 2 | |
| Deletion | Chr4 | 15984135 | 15984141 | 6 | |
| Deletion | Chr2 | 17801431 | 17801432 | 1 | |
| Deletion | Chr9 | 18968001 | 18980000 | 11999 | 2 |
| Deletion | Chr2 | 19750088 | 19750096 | 8 | |
| Deletion | Chr11 | 24945103 | 24945107 | 4 | |
| Deletion | Chr1 | 35128911 | 35128912 | 1 |
Insertions: 5
Inversions: 3
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr8 | 1361702 | 1362232 | |
| Inversion | Chr2 | 20792881 | 21306648 | LOC_Os02g34660 |
| Inversion | Chr2 | 20792889 | 21306656 | LOC_Os02g34660 |
Translocations: 5
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr4 | 7677812 | Chr1 | 19525915 | LOC_Os04g13770 |
| Translocation | Chr6 | 9741808 | Chr5 | 18325130 | |
| Translocation | Chr8 | 10828772 | Chr4 | 21079421 | |
| Translocation | Chr10 | 15534737 | Chr7 | 7220331 | |
| Translocation | Chr3 | 26496232 | Chr2 | 5969105 |
