Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN2765-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN2765-S Alignment File |
| Seed Availability | No |
| Mapping (Hover to Zoom-In) | 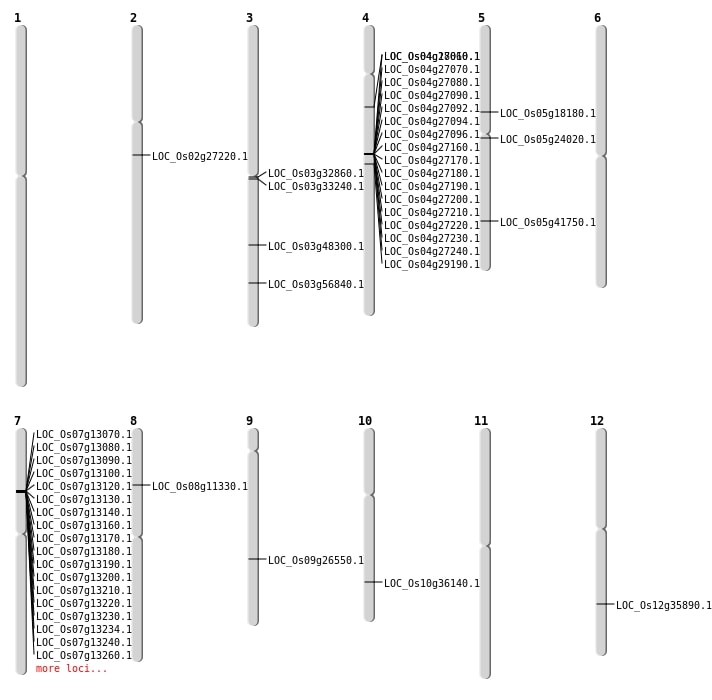
|
| Variant Information |
|---|
Single base substitutions: 82
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 11922588 | C-T | HET | ||
| SBS | Chr1 | 12893504 | A-G | HET | ||
| SBS | Chr1 | 14016053 | T-A | Homo | ||
| SBS | Chr1 | 19283800 | T-C | HET | ||
| SBS | Chr1 | 25577722 | G-T | HET | ||
| SBS | Chr1 | 29288572 | A-T | HET | ||
| SBS | Chr1 | 31636038 | T-G | HET | ||
| SBS | Chr1 | 36263939 | A-G | HET | ||
| SBS | Chr1 | 40115858 | C-T | HET | ||
| SBS | Chr10 | 1852779 | T-A | HET | ||
| SBS | Chr10 | 19318754 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os10g36140 |
| SBS | Chr10 | 2289958 | T-C | HET | ||
| SBS | Chr10 | 9193666 | G-A | HET | ||
| SBS | Chr10 | 9823601 | G-T | HET | ||
| SBS | Chr11 | 11199616 | C-T | HET | ||
| SBS | Chr11 | 13850170 | C-A | Homo | ||
| SBS | Chr11 | 23816458 | G-A | Homo | ||
| SBS | Chr11 | 26115239 | T-C | HET | ||
| SBS | Chr11 | 26501494 | A-G | HET | ||
| SBS | Chr11 | 2740387 | C-T | HET | ||
| SBS | Chr11 | 7969526 | G-A | HET | ||
| SBS | Chr11 | 8242108 | C-A | HET | ||
| SBS | Chr11 | 9258824 | G-T | HET | ||
| SBS | Chr11 | 975856 | C-T | HET | ||
| SBS | Chr12 | 10594129 | C-A | HET | ||
| SBS | Chr12 | 10908092 | A-G | HET | ||
| SBS | Chr12 | 11210440 | C-T | HET | ||
| SBS | Chr12 | 12659043 | G-C | Homo | ||
| SBS | Chr12 | 12818534 | C-A | HET | ||
| SBS | Chr12 | 13278861 | T-A | HET | ||
| SBS | Chr12 | 13278862 | G-T | HET | ||
| SBS | Chr12 | 14551657 | T-A | Homo | ||
| SBS | Chr12 | 18063017 | G-T | HET | ||
| SBS | Chr12 | 21871123 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os12g35890 |
| SBS | Chr12 | 25259106 | G-A | HET | ||
| SBS | Chr12 | 322860 | C-A | HET | ||
| SBS | Chr12 | 7670314 | C-T | HET | ||
| SBS | Chr2 | 16013916 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os02g27220 |
| SBS | Chr2 | 17703887 | C-A | HET | ||
| SBS | Chr2 | 224636 | G-T | HET | ||
| SBS | Chr2 | 3948118 | A-T | HET | ||
| SBS | Chr2 | 6747108 | A-T | Homo | ||
| SBS | Chr3 | 18803329 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os03g32860 |
| SBS | Chr3 | 19020070 | C-T | Homo | NON_SYNONYMOUS_CODING | LOC_Os03g33240 |
| SBS | Chr3 | 2746316 | G-T | HET | ||
| SBS | Chr3 | 31234634 | C-T | Homo | ||
| SBS | Chr3 | 32387051 | A-C | Homo | NON_SYNONYMOUS_CODING | LOC_Os03g56840 |
| SBS | Chr3 | 350897 | G-A | HET | ||
| SBS | Chr3 | 9303660 | C-T | Homo | ||
| SBS | Chr4 | 12765882 | C-T | HET | ||
| SBS | Chr4 | 15044776 | T-A | HET | ||
| SBS | Chr4 | 17307235 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os04g29190 |
| SBS | Chr4 | 18404959 | G-A | HET | ||
| SBS | Chr5 | 10447889 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os05g18180 |
| SBS | Chr5 | 12997757 | G-A | HET | ||
| SBS | Chr5 | 16088682 | G-A | HET | ||
| SBS | Chr5 | 21535003 | G-T | Homo | ||
| SBS | Chr5 | 530908 | T-A | HET | ||
| SBS | Chr5 | 6617047 | C-A | HET | ||
| SBS | Chr5 | 743742 | A-T | Homo | ||
| SBS | Chr6 | 11941646 | C-T | HET | ||
| SBS | Chr6 | 12039239 | T-A | HET | ||
| SBS | Chr6 | 12805544 | G-A | HET | ||
| SBS | Chr6 | 13005025 | T-C | HET | ||
| SBS | Chr6 | 13596907 | G-A | HET | ||
| SBS | Chr6 | 15301990 | G-A | HET | ||
| SBS | Chr6 | 16137509 | G-A | HET | ||
| SBS | Chr6 | 16836877 | G-T | HET | ||
| SBS | Chr6 | 17192406 | G-A | HET | ||
| SBS | Chr6 | 22233198 | C-A | HET | ||
| SBS | Chr7 | 14135240 | G-T | HET | ||
| SBS | Chr7 | 16135349 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os07g27630 |
| SBS | Chr7 | 209962 | G-T | HET | ||
| SBS | Chr7 | 21201885 | G-C | HET | ||
| SBS | Chr7 | 8984277 | C-T | HET | ||
| SBS | Chr7 | 9262232 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os07g15940 |
| SBS | Chr8 | 18700711 | T-G | Homo | ||
| SBS | Chr8 | 21153905 | T-A | HET | ||
| SBS | Chr8 | 6662149 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os08g11330 |
| SBS | Chr8 | 9456123 | T-C | HET | ||
| SBS | Chr8 | 9456156 | G-T | HET | ||
| SBS | Chr9 | 16051747 | C-G | HET | NON_SYNONYMOUS_CODING | LOC_Os09g26550 |
Deletions: 16
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr8 | 911975 | 911976 | 1 | |
| Deletion | Chr12 | 2506649 | 2506650 | 1 | |
| Deletion | Chr7 | 2876231 | 2876252 | 21 | |
| Deletion | Chr2 | 3242883 | 3242886 | 3 | |
| Deletion | Chr5 | 5615782 | 5615788 | 6 | |
| Deletion | Chr7 | 7491001 | 7627000 | 135999 | 19 |
| Deletion | Chr1 | 7910190 | 7910191 | 1 | |
| Deletion | Chr8 | 12169966 | 12169967 | 1 | |
| Deletion | Chr6 | 12630193 | 12630195 | 2 | |
| Deletion | Chr11 | 15501827 | 15501831 | 4 | |
| Deletion | Chr4 | 16011001 | 16020000 | 8999 | 3 |
| Deletion | Chr4 | 16023001 | 16097000 | 73999 | 12 |
| Deletion | Chr9 | 17406401 | 17406403 | 2 | |
| Deletion | Chr11 | 25027051 | 25027052 | 1 | |
| Deletion | Chr3 | 27491001 | 27501000 | 9999 | LOC_Os03g48300 |
| Deletion | Chr8 | 27908213 | 27908214 | 1 |
Insertions: 5
Inversions: 6
Translocations: 6
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr8 | 4116725 | Chr5 | 24425573 | 2 |
| Translocation | Chr8 | 4116952 | Chr5 | 14470343 | |
| Translocation | Chr12 | 4332524 | Chr1 | 25576631 | |
| Translocation | Chr5 | 14470339 | Chr4 | 16004977 | 2 |
| Translocation | Chr5 | 24425574 | Chr4 | 16004977 | 2 |
| Translocation | Chr12 | 27094165 | Chr4 | 9895923 | 2 |
