Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN2790-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN2790-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 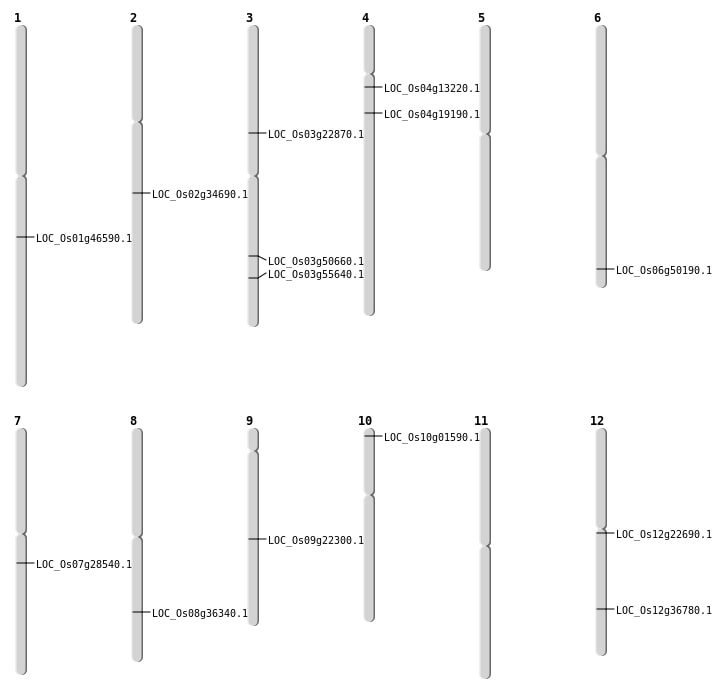
|
| Variant Information |
|---|
Single base substitutions: 39
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 20026652 | C-A | Homo | ||
| SBS | Chr1 | 34919998 | G-C | HET | ||
| SBS | Chr1 | 39833413 | C-T | HET | ||
| SBS | Chr1 | 9208525 | G-A | Homo | ||
| SBS | Chr10 | 15942131 | C-T | HET | ||
| SBS | Chr10 | 22649473 | A-G | Homo | ||
| SBS | Chr11 | 11395989 | T-A | Homo | ||
| SBS | Chr11 | 13925178 | G-T | Homo | ||
| SBS | Chr11 | 14918136 | C-T | HET | ||
| SBS | Chr11 | 26337882 | C-A | HET | ||
| SBS | Chr11 | 4751276 | G-C | Homo | ||
| SBS | Chr12 | 12810644 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os12g22690 |
| SBS | Chr12 | 22554428 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os12g36780 |
| SBS | Chr12 | 8028466 | T-C | HET | ||
| SBS | Chr3 | 22907941 | A-T | HET | ||
| SBS | Chr3 | 24511142 | T-A | HET | ||
| SBS | Chr3 | 32893619 | G-C | Homo | ||
| SBS | Chr3 | 5098253 | C-T | Homo | ||
| SBS | Chr4 | 10657466 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os04g19190 |
| SBS | Chr4 | 7324996 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os04g13220 |
| SBS | Chr5 | 14268821 | C-T | HET | ||
| SBS | Chr5 | 19648367 | C-G | HET | ||
| SBS | Chr5 | 5038012 | G-A | HET | ||
| SBS | Chr5 | 9424646 | C-T | HET | ||
| SBS | Chr6 | 15753470 | G-A | HET | ||
| SBS | Chr6 | 30388203 | C-T | Homo | NON_SYNONYMOUS_CODING | LOC_Os06g50190 |
| SBS | Chr6 | 31124355 | G-A | Homo | ||
| SBS | Chr6 | 6908640 | G-A | HET | ||
| SBS | Chr7 | 16699783 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os07g28540 |
| SBS | Chr7 | 17180513 | G-A | HET | ||
| SBS | Chr7 | 18104029 | A-G | HET | ||
| SBS | Chr7 | 19719253 | C-T | HET | ||
| SBS | Chr7 | 21251409 | A-G | HET | ||
| SBS | Chr7 | 6709455 | A-G | HET | ||
| SBS | Chr8 | 21720002 | C-G | HET | ||
| SBS | Chr8 | 26572554 | C-A | HET | ||
| SBS | Chr9 | 18362967 | C-T | Homo | ||
| SBS | Chr9 | 5839359 | G-A | HET | ||
| SBS | Chr9 | 9369421 | G-A | HET |
Deletions: 23
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr3 | 1937366 | 1937377 | 11 | |
| Deletion | Chr8 | 4175729 | 4175734 | 5 | |
| Deletion | Chr6 | 4304988 | 4304990 | 2 | |
| Deletion | Chr6 | 5981612 | 5981614 | 2 | |
| Deletion | Chr12 | 8166198 | 8166202 | 4 | |
| Deletion | Chr7 | 9593394 | 9593395 | 1 | |
| Deletion | Chr9 | 9610829 | 9610830 | 1 | |
| Deletion | Chr9 | 9611124 | 9611137 | 13 | |
| Deletion | Chr3 | 13220853 | 13220855 | 2 | LOC_Os03g22870 |
| Deletion | Chr9 | 13475262 | 13475266 | 4 | LOC_Os09g22300 |
| Deletion | Chr6 | 13707370 | 13707377 | 7 | |
| Deletion | Chr11 | 13772370 | 13772377 | 7 | |
| Deletion | Chr10 | 14990856 | 14990857 | 1 | |
| Deletion | Chr2 | 15882418 | 15882423 | 5 | |
| Deletion | Chr12 | 17891911 | 17891917 | 6 | |
| Deletion | Chr10 | 19377101 | 19377115 | 14 | |
| Deletion | Chr2 | 20814655 | 20814657 | 2 | LOC_Os02g34690 |
| Deletion | Chr9 | 21220488 | 21220496 | 8 | |
| Deletion | Chr8 | 22907633 | 22907639 | 6 | LOC_Os08g36340 |
| Deletion | Chr7 | 25661139 | 25661141 | 2 | |
| Deletion | Chr1 | 26524193 | 26524217 | 24 | LOC_Os01g46590 |
| Deletion | Chr1 | 31510386 | 31510392 | 6 | |
| Deletion | Chr3 | 36067592 | 36067595 | 3 |
Insertions: 1
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr3 | 28928359 | 28928361 | 3 | LOC_Os03g50660 |
No Inversion
Translocations: 3
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr10 | 390825 | Chr5 | 10086180 | LOC_Os10g01590 |
| Translocation | Chr11 | 9540999 | Chr3 | 31682825 | 2 |
| Translocation | Chr11 | 9541127 | Chr3 | 31680040 |
