Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN3659-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN3659-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 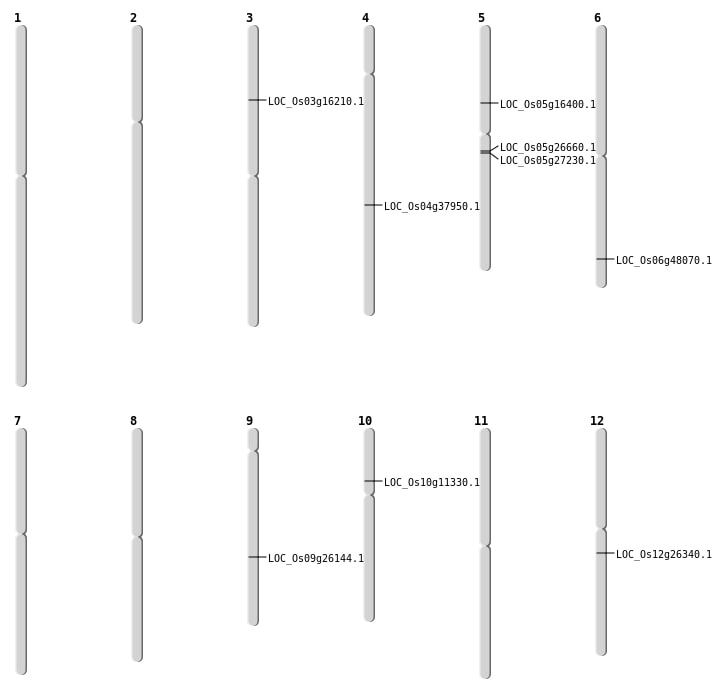
|
| Variant Information |
|---|
Single base substitutions: 47
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 10491263 | G-A | HET | ||
| SBS | Chr1 | 11081936 | T-G | HET | ||
| SBS | Chr1 | 15327594 | G-A | HET | ||
| SBS | Chr1 | 17924307 | G-A | HET | ||
| SBS | Chr10 | 19258197 | G-T | HET | ||
| SBS | Chr10 | 3959020 | G-A | HET | ||
| SBS | Chr10 | 6285966 | G-T | Homo | NON_SYNONYMOUS_CODING | LOC_Os10g11330 |
| SBS | Chr10 | 6966343 | G-A | Homo | ||
| SBS | Chr11 | 15180057 | G-A | HET | ||
| SBS | Chr11 | 2258590 | A-C | HET | ||
| SBS | Chr11 | 2326337 | C-T | HET | ||
| SBS | Chr11 | 7587452 | G-A | HET | ||
| SBS | Chr11 | 7753063 | A-T | HET | ||
| SBS | Chr11 | 8135402 | G-T | HET | ||
| SBS | Chr12 | 11756978 | G-A | HET | ||
| SBS | Chr12 | 15387881 | G-C | HET | NON_SYNONYMOUS_CODING | LOC_Os12g26340 |
| SBS | Chr12 | 16834899 | G-A | HET | ||
| SBS | Chr2 | 21012866 | G-A | HET | ||
| SBS | Chr2 | 2253004 | G-T | Homo | ||
| SBS | Chr2 | 30077003 | C-T | HET | ||
| SBS | Chr3 | 12843807 | C-A | HET | ||
| SBS | Chr3 | 15609327 | G-A | HET | ||
| SBS | Chr3 | 15921457 | C-T | HET | ||
| SBS | Chr3 | 15976152 | A-G | HET | ||
| SBS | Chr3 | 16332736 | C-T | HET | ||
| SBS | Chr3 | 18584766 | A-G | HET | ||
| SBS | Chr3 | 22420424 | T-C | HET | ||
| SBS | Chr3 | 27940733 | G-A | HET | ||
| SBS | Chr3 | 8941166 | T-A | HET | NON_SYNONYMOUS_CODING | LOC_Os03g16210 |
| SBS | Chr4 | 1338487 | C-A | HET | ||
| SBS | Chr4 | 21191509 | G-A | HET | ||
| SBS | Chr4 | 22568892 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os04g37950 |
| SBS | Chr4 | 34087049 | C-T | HET | ||
| SBS | Chr5 | 14746869 | T-C | HET | ||
| SBS | Chr5 | 19487017 | A-T | HET | ||
| SBS | Chr5 | 9301111 | T-C | HET | NON_SYNONYMOUS_CODING | LOC_Os05g16400 |
| SBS | Chr6 | 2421030 | C-T | Homo | ||
| SBS | Chr6 | 28564988 | G-A | HET | ||
| SBS | Chr6 | 5654750 | G-A | HET | ||
| SBS | Chr7 | 19840866 | T-G | HET | ||
| SBS | Chr7 | 22632337 | T-G | HET | ||
| SBS | Chr8 | 26612928 | C-T | HET | ||
| SBS | Chr8 | 27316599 | A-G | HET | ||
| SBS | Chr8 | 3872389 | G-A | HET | ||
| SBS | Chr8 | 6694871 | G-C | HET | ||
| SBS | Chr8 | 7803629 | A-G | HET | ||
| SBS | Chr9 | 20504493 | G-A | HET |
Deletions: 10
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr5 | 2616589 | 2616594 | 5 | |
| Deletion | Chr5 | 4903222 | 4903233 | 11 | |
| Deletion | Chr2 | 6270143 | 6270144 | 1 | |
| Deletion | Chr9 | 8090729 | 8090734 | 5 | |
| Deletion | Chr2 | 8443156 | 8443159 | 3 | |
| Deletion | Chr3 | 8863551 | 8863563 | 12 | |
| Deletion | Chr10 | 13154250 | 13154262 | 12 | |
| Deletion | Chr9 | 15761707 | 15761712 | 5 | LOC_Os09g26144 |
| Deletion | Chr2 | 20170404 | 20170420 | 16 | |
| Deletion | Chr4 | 20516185 | 20516187 | 2 |
Insertions: 9
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr1 | 23067221 | 23067224 | 4 | |
| Insertion | Chr10 | 14051554 | 14051555 | 2 | |
| Insertion | Chr11 | 12095254 | 12095256 | 3 | |
| Insertion | Chr11 | 15036274 | 15036274 | 1 | |
| Insertion | Chr12 | 24258687 | 24258688 | 2 | |
| Insertion | Chr2 | 16230649 | 16230649 | 1 | |
| Insertion | Chr5 | 9941976 | 9941976 | 1 | |
| Insertion | Chr6 | 29086294 | 29086294 | 1 | LOC_Os06g48070 |
| Insertion | Chr8 | 11819534 | 11819534 | 1 |
Inversions: 2
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr5 | 15482600 | 15833809 | 2 |
| Inversion | Chr6 | 22483312 | 22521117 |
Translocations: 3
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr12 | 730999 | Chr11 | 690967 | |
| Translocation | Chr11 | 15630258 | Chr9 | 3034010 | |
| Translocation | Chr11 | 24337241 | Chr2 | 3897894 |
