Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN813-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN813-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 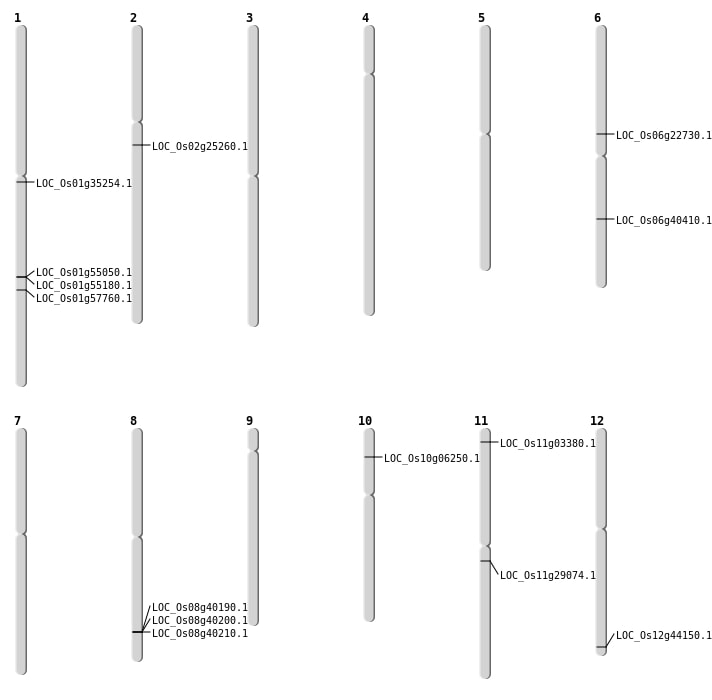
|
| Variant Information |
|---|
Single base substitutions: 23
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 19518787 | T-C | HET | NON_SYNONYMOUS_CODING | LOC_Os01g35254 |
| SBS | Chr1 | 24413184 | C-T | HET | INTERGENIC | |
| SBS | Chr1 | 31747376 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os01g55180 |
| SBS | Chr1 | 33404613 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os01g57760 |
| SBS | Chr1 | 3973512 | C-T | HET | INTERGENIC | |
| SBS | Chr10 | 17247055 | C-A | HET | INTERGENIC | |
| SBS | Chr10 | 3207169 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os10g06250 |
| SBS | Chr11 | 1272055 | T-C | HOMO | INTERGENIC | |
| SBS | Chr12 | 15132237 | T-G | HOMO | INTRON | |
| SBS | Chr12 | 26049422 | A-T | HET | INTERGENIC | |
| SBS | Chr12 | 4007031 | C-T | HOMO | INTRON | |
| SBS | Chr2 | 14710557 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os02g25260 |
| SBS | Chr2 | 4194148 | T-C | HET | INTERGENIC | |
| SBS | Chr3 | 34738761 | A-G | HET | INTERGENIC | |
| SBS | Chr4 | 6296224 | C-G | HET | INTRON | |
| SBS | Chr6 | 13220265 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os06g22730 |
| SBS | Chr6 | 22129511 | G-A | HET | INTERGENIC | |
| SBS | Chr6 | 4372625 | G-A | HET | INTRON | |
| SBS | Chr7 | 14458091 | C-T | HOMO | INTRON | |
| SBS | Chr8 | 4288780 | T-C | HET | INTERGENIC | |
| SBS | Chr9 | 10050665 | G-A | HET | SYNONYMOUS_CODING | |
| SBS | Chr9 | 10310495 | C-T | HOMO | INTERGENIC | |
| SBS | Chr9 | 1876004 | C-A | HOMO | INTERGENIC |
Deletions: 19
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr4 | 417140 | 417142 | 2 | |
| Deletion | Chr11 | 1274821 | 1274829 | 8 | LOC_Os11g03380 |
| Deletion | Chr8 | 2335148 | 2335158 | 10 | |
| Deletion | Chr3 | 3555381 | 3555390 | 9 | |
| Deletion | Chr1 | 3910333 | 3910339 | 6 | |
| Deletion | Chr5 | 4715514 | 4715523 | 9 | |
| Deletion | Chr11 | 5201873 | 5201882 | 9 | |
| Deletion | Chr6 | 7628156 | 7628219 | 63 | |
| Deletion | Chr12 | 11266833 | 11266838 | 5 | |
| Deletion | Chr11 | 11953558 | 11953560 | 2 | |
| Deletion | Chr10 | 16357583 | 16357590 | 7 | |
| Deletion | Chr11 | 16843197 | 16843198 | 1 | LOC_Os11g29074 |
| Deletion | Chr5 | 17035141 | 17035150 | 9 | |
| Deletion | Chr11 | 21155777 | 21155800 | 23 | |
| Deletion | Chr6 | 24063552 | 24063558 | 6 | LOC_Os06g40410 |
| Deletion | Chr8 | 25436281 | 25442987 | 6706 | 3 |
| Deletion | Chr6 | 28850963 | 28850983 | 20 | |
| Deletion | Chr1 | 31655240 | 31655526 | 286 | |
| Deletion | Chr1 | 35361699 | 35361711 | 12 |
Insertions: 5
No Inversion
Translocations: 4
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr9 | 14842568 | Chr1 | 31655516 | LOC_Os01g55050 |
| Translocation | Chr9 | 14842578 | Chr1 | 31655246 | LOC_Os01g55050 |
| Translocation | Chr12 | 27391441 | Chr8 | 25442988 | LOC_Os12g44150 |
| Translocation | Chr12 | 27391450 | Chr8 | 25436457 | LOC_Os12g44150 |
