Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN893-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN893-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 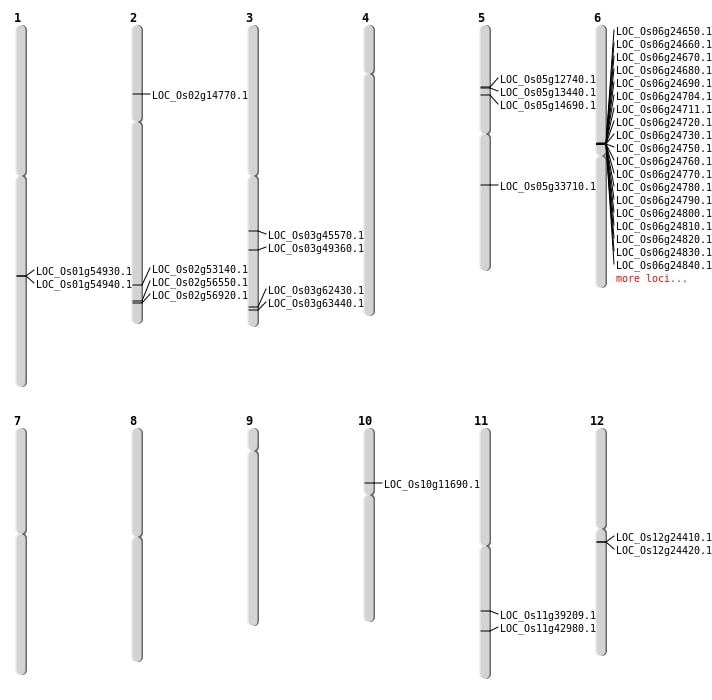
|
| Variant Information |
|---|
Single base substitutions: 30
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 18737321 | C-G | HET | INTRON | |
| SBS | Chr1 | 22081116 | G-A | HET | INTERGENIC | |
| SBS | Chr1 | 248081 | G-A | HET | INTERGENIC | |
| SBS | Chr1 | 27178666 | A-G | HET | INTERGENIC | |
| SBS | Chr1 | 30472649 | C-T | HOMO | UTR_5_PRIME | |
| SBS | Chr1 | 36323420 | G-A | HET | INTERGENIC | |
| SBS | Chr10 | 6457271 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os10g11690 |
| SBS | Chr11 | 20117669 | G-A | HET | INTERGENIC | |
| SBS | Chr11 | 25907133 | G-C | HET | NON_SYNONYMOUS_CODING | LOC_Os11g42980 |
| SBS | Chr11 | 4154426 | A-T | HOMO | INTRON | |
| SBS | Chr11 | 7387628 | C-T | HET | INTERGENIC | |
| SBS | Chr12 | 24613480 | C-T | HET | INTERGENIC | |
| SBS | Chr2 | 14658172 | G-C | HET | INTERGENIC | |
| SBS | Chr2 | 6760624 | G-A | HET | INTERGENIC | |
| SBS | Chr2 | 7320621 | C-T | HET | INTERGENIC | |
| SBS | Chr3 | 14898605 | G-A | HET | INTERGENIC | |
| SBS | Chr3 | 17129522 | T-G | HET | INTERGENIC | |
| SBS | Chr3 | 32745092 | G-A | HET | UTR_5_PRIME | |
| SBS | Chr4 | 1304862 | C-G | HET | INTERGENIC | |
| SBS | Chr5 | 1597388 | G-T | HET | INTERGENIC | |
| SBS | Chr5 | 19842763 | T-C | HET | NON_SYNONYMOUS_CODING | LOC_Os05g33710 |
| SBS | Chr5 | 8353133 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os05g14690 |
| SBS | Chr7 | 11789226 | A-T | HOMO | SYNONYMOUS_CODING | |
| SBS | Chr7 | 9668637 | C-T | HOMO | INTERGENIC | |
| SBS | Chr8 | 12229231 | C-T | HOMO | INTERGENIC | |
| SBS | Chr8 | 14031356 | C-T | HET | SYNONYMOUS_CODING | |
| SBS | Chr8 | 5119919 | G-A | HET | INTRON | |
| SBS | Chr9 | 1400581 | G-C | HET | INTERGENIC | |
| SBS | Chr9 | 20804447 | A-C | HOMO | INTERGENIC | |
| SBS | Chr9 | 4034119 | T-A | HET | INTRON |
Deletions: 20
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr7 | 5772707 | 5772717 | 10 | |
| Deletion | Chr2 | 6755313 | 6755319 | 6 | |
| Deletion | Chr5 | 6941513 | 6941518 | 5 | |
| Deletion | Chr11 | 11045695 | 11045711 | 16 | |
| Deletion | Chr12 | 13924001 | 13938000 | 14000 | 2 |
| Deletion | Chr6 | 14470001 | 14639000 | 169000 | 29 |
| Deletion | Chr8 | 18612882 | 18612887 | 5 | |
| Deletion | Chr6 | 19723001 | 19930000 | 207000 | 28 |
| Deletion | Chr11 | 23348213 | 23348215 | 2 | LOC_Os11g39209 |
| Deletion | Chr3 | 25719758 | 25719762 | 4 | LOC_Os03g45570 |
| Deletion | Chr1 | 25859067 | 25859072 | 5 | |
| Deletion | Chr5 | 26337843 | 26340791 | 2948 | |
| Deletion | Chr1 | 31052818 | 31052827 | 9 | |
| Deletion | Chr1 | 31596010 | 31601553 | 5543 | 2 |
| Deletion | Chr2 | 34554219 | 34554237 | 18 | |
| Deletion | Chr2 | 34640877 | 34640898 | 21 | LOC_Os02g56550 |
| Deletion | Chr2 | 34890157 | 34890177 | 20 | LOC_Os02g56920 |
| Deletion | Chr3 | 35362942 | 35362945 | 3 | LOC_Os03g62430 |
| Deletion | Chr3 | 35841700 | 35841703 | 3 | LOC_Os03g63440 |
| Deletion | Chr1 | 37465903 | 37465906 | 3 |
Insertions: 1
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr2 | 9452252 | 9452252 | 1 |
Inversions: 5
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr5 | 7310719 | 7456455 | 2 |
| Inversion | Chr5 | 7310742 | 7456471 | 2 |
| Inversion | Chr2 | 8090962 | 8183885 | LOC_Os02g14770 |
| Inversion | Chr2 | 8091006 | 8183862 | LOC_Os02g14770 |
| Inversion | Chr2 | 31540754 | 32530921 | LOC_Os02g53140 |
Translocations: 2
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr3 | 28096154 | Chr2 | 34504967 | LOC_Os03g49360 |
| Translocation | Chr3 | 28097772 | Chr2 | 34505021 | LOC_Os03g49360 |
