Search Nipponbare
| Mutant Information |
|---|
| Mutant Name | FN920-S [Download] |
| Generation | M2 |
| Genus Species | Oryza Sativa |
| Cultivar | Kitaake |
| Alignment File (BAM File) | Download FN920-S Alignment File |
| Seed Availability | Yes [Order Seeds] |
| Mapping (Hover to Zoom-In) | 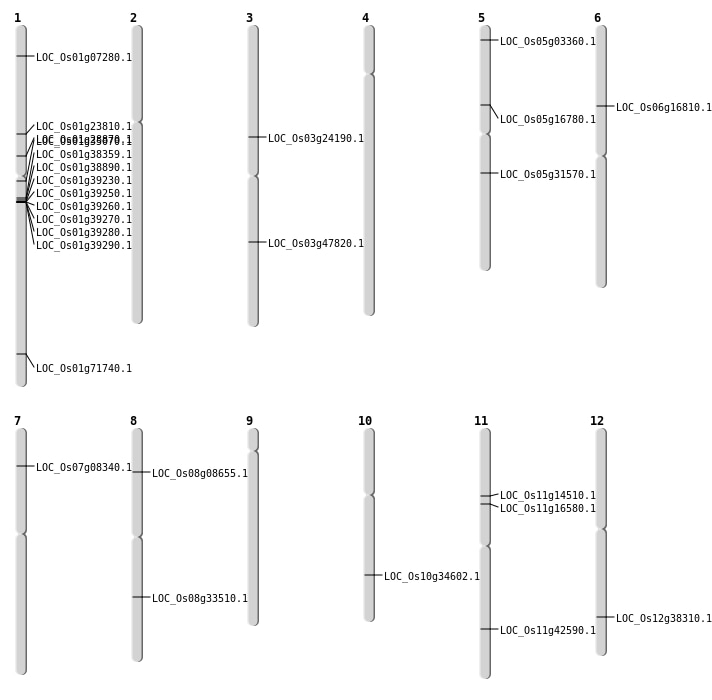
|
| Variant Information |
|---|
Single base substitutions: 33
| Variant Type | Chromosome | Position | Change | Genotype | Effect | Gene Affected |
|---|---|---|---|---|---|---|
| SBS | Chr1 | 13396404 | C-G | HET | NON_SYNONYMOUS_CODING | LOC_Os01g23810 |
| SBS | Chr1 | 14258265 | T-G | HET | INTERGENIC | |
| SBS | Chr1 | 16165140 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os01g28870 |
| SBS | Chr1 | 16165142 | A-G | HET | NON_SYNONYMOUS_CODING | LOC_Os01g28870 |
| SBS | Chr1 | 19418803 | C-T | HET | NON_SYNONYMOUS_CODING | LOC_Os01g35070 |
| SBS | Chr10 | 22721671 | G-C | HET | INTERGENIC | |
| SBS | Chr11 | 22918438 | C-T | HET | INTERGENIC | |
| SBS | Chr11 | 24923574 | C-T | HET | INTERGENIC | |
| SBS | Chr11 | 25005680 | C-T | HET | UTR_3_PRIME | |
| SBS | Chr12 | 11445947 | C-T | HET | INTERGENIC | |
| SBS | Chr12 | 21524734 | T-C | HET | INTERGENIC | |
| SBS | Chr12 | 8600264 | A-G | HET | INTRON | |
| SBS | Chr3 | 13752527 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os03g24190 |
| SBS | Chr3 | 27161284 | C-A | HET | NON_SYNONYMOUS_CODING | LOC_Os03g47820 |
| SBS | Chr4 | 31808696 | G-T | HET | INTERGENIC | |
| SBS | Chr5 | 1378163 | C-T | HET | STOP_GAINED | LOC_Os05g03360 |
| SBS | Chr5 | 16154981 | A-T | HOMO | INTERGENIC | |
| SBS | Chr5 | 18380786 | C-T | HOMO | SYNONYMOUS_CODING | |
| SBS | Chr5 | 18380787 | C-T | HOMO | STOP_GAINED | LOC_Os05g31570 |
| SBS | Chr5 | 25870077 | T-A | HOMO | INTERGENIC | |
| SBS | Chr5 | 9553608 | T-C | HOMO | NON_SYNONYMOUS_CODING | LOC_Os05g16780 |
| SBS | Chr6 | 22532586 | G-A | HET | INTRON | |
| SBS | Chr7 | 17952028 | A-G | HET | INTERGENIC | |
| SBS | Chr7 | 20110813 | A-T | HET | INTERGENIC | |
| SBS | Chr7 | 28167603 | A-G | HET | SYNONYMOUS_CODING | |
| SBS | Chr7 | 4273956 | A-G | HET | INTERGENIC | |
| SBS | Chr7 | 4273958 | C-G | HET | INTERGENIC | |
| SBS | Chr8 | 15488814 | T-G | HOMO | INTERGENIC | |
| SBS | Chr8 | 20914925 | G-T | HET | NON_SYNONYMOUS_CODING | LOC_Os08g33510 |
| SBS | Chr8 | 23950530 | C-T | HET | INTERGENIC | |
| SBS | Chr8 | 5011929 | G-A | HET | NON_SYNONYMOUS_CODING | LOC_Os08g08655 |
| SBS | Chr9 | 4621560 | A-G | HOMO | INTERGENIC | |
| SBS | Chr9 | 5007943 | T-C | HET | INTRON |
Deletions: 17
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Deletion | Chr10 | 918885 | 918888 | 3 | |
| Deletion | Chr7 | 7341299 | 7341524 | 225 | |
| Deletion | Chr2 | 8582364 | 8582365 | 1 | |
| Deletion | Chr11 | 9030172 | 9030177 | 5 | |
| Deletion | Chr6 | 9735365 | 9735366 | 1 | LOC_Os06g16810 |
| Deletion | Chr7 | 16927049 | 16927055 | 6 | |
| Deletion | Chr7 | 16962107 | 16962108 | 1 | |
| Deletion | Chr6 | 17499979 | 17499982 | 3 | |
| Deletion | Chr8 | 21032874 | 21032889 | 15 | |
| Deletion | Chr7 | 21095157 | 21095163 | 6 | |
| Deletion | Chr12 | 22063382 | 22063388 | 6 | |
| Deletion | Chr1 | 22066001 | 22116000 | 50000 | 6 |
| Deletion | Chr4 | 22738914 | 22740740 | 1826 | |
| Deletion | Chr2 | 23544024 | 23544025 | 1 | |
| Deletion | Chr11 | 28833662 | 28833663 | 1 | |
| Deletion | Chr2 | 30484808 | 30484811 | 3 | |
| Deletion | Chr1 | 42046681 | 42046685 | 4 |
Insertions: 3
| Variant Type | Chromosome | Start | End | Size (bp) | Number of Genes |
|---|---|---|---|---|---|
| Insertion | Chr11 | 19595026 | 19595027 | 2 | |
| Insertion | Chr2 | 8103302 | 8103305 | 4 | |
| Insertion | Chr7 | 26505874 | 26505875 | 2 |
Inversions: 6
| Variant Type | Chromosome | Position 1 | Position 2 | Number of Genes |
|---|---|---|---|---|
| Inversion | Chr11 | 8151093 | 9188429 | 2 |
| Inversion | Chr1 | 21547555 | 22001960 | LOC_Os01g38359 |
| Inversion | Chr1 | 21823922 | 22001936 | LOC_Os01g38890 |
| Inversion | Chr1 | 22014763 | 22585767 | |
| Inversion | Chr1 | 41565481 | 41774910 | LOC_Os01g71740 |
| Inversion | Chr1 | 41565564 | 41774636 | LOC_Os01g71740 |
Translocations: 9
| Variant Type | Chromosome | Position 1 | Chromosome | Position 2 | Number of Genes |
|---|---|---|---|---|---|
| Translocation | Chr7 | 4269481 | Chr1 | 21884886 | LOC_Os07g08340 |
| Translocation | Chr7 | 4270192 | Chr1 | 22014752 | LOC_Os07g08340 |
| Translocation | Chr10 | 19503141 | Chr7 | 4163450 | |
| Translocation | Chr10 | 19503214 | Chr7 | 4219012 | |
| Translocation | Chr9 | 19842088 | Chr1 | 3432303 | LOC_Os01g07280 |
| Translocation | Chr12 | 23517505 | Chr7 | 4270193 | 2 |
| Translocation | Chr12 | 23517508 | Chr7 | 4276754 | LOC_Os12g38310 |
| Translocation | Chr11 | 25637609 | Chr7 | 4163382 | LOC_Os11g42590 |
| Translocation | Chr11 | 25638072 | Chr10 | 18448165 | 2 |
